DMR 104 chr1:10509901-10510100 (200bp)
| DMR Stats | |
|---|---|
| Position | chr1:10509901-10510100 (View on UCSC) |
| Width | 200bp |
| Genomic Context | promoter |
| Nearest Genes | APITD1-CORT, CORT (0bp away) |
| ANODEV p-value | 0.0253035283924 |
| Frequencies | Benign: 5.0 (71.43%), Low: 6.0 (100.0%), High: 8.0 (88.89 %) |
| Frequent? |
Region Count Boxplot

Region Overview

Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 104 | APITD1-CORT | 197 | uc001arf.3 | chr1 | 10490159 | 10512060 | + | 200 | 100.0 | 0.55 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (100), E5 (0) | 0.0 |
| 104 | CORT | 197 | uc001ari.4 | chr1 | 10509776 | 10512060 | + | 200 | 100.0 | 0.25 | 0 | 100.0 | E1 (100), I1 (0), E2 (0) | 0.0 |
| 104 | APITD1-CORT | 197 | uc021ogd.1 | chr1 | 10490159 | 10512060 | + | 200 | 100.0 | 0.18 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (100), E4 (0) | 0.0 |
| 104 | APITD1-CORT | 197 | uc021ogf.1 | chr1 | 10490804 | 10512060 | + | 200 | 100.0 | 0.21 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (100), E4 (0) | 0.0 |
| 104 | APITD1-CORT | 197 | uc021ogg.1 | chr1 | 10490804 | 10512060 | + | 200 | 100.0 | 0.28 | 1 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (100), E3 (0) | 0.0 |
| 104 | APITD1-CORT | 197 | uc031ple.1 | chr1 | 10490159 | 10512060 | + | 200 | 100.0 | 0.28 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (100), E6 (0) | 0.0 |
- 6 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 104 | chr1 | 10509981 | 10511035 | wgEncodeRegDnaseClusteredV2 | 28 | score:413, srow:8459 |
| 104 | chr1 | 10509906 | 10509919 | tfbsConsSites | V$ELK1_02 | score:867, zScore:1.74, srow:19617 |
| 104 | chr1 | 10509907 | 10509916 | tfbsConsSites | V$CETS1P54_01 | score:946, zScore:1.83, srow:19618 |
| 104 | chr1 | 10510025 | 10510032 | tfbsConsSites | V$NKX25_02 | score:947, zScore:2.34, srow:19619 |
| 104 | chr1 | 10510083 | 10510093 | tfbsConsSites | V$AP1FJ_Q2 | score:929, zScore:2.09, srow:19620 |
| 104 | chr1 | 10510083 | 10510097 | tfbsConsSites | V$TAXCREB_01 | score:834, zScore:2.71, srow:19621 |
| 104 | chr1 | 10509909 | 10509915 | phastConsElements100way | lod=33 | score:340, srow:38621 |
| 104 | chr1 | 10509936 | 10509940 | phastConsElements100way | lod=23 | score:304, srow:38622 |
| 104 | chr1 | 10509978 | 10509988 | phastConsElements100way | lod=22 | score:300, srow:38623 |
| 104 | chr1 | 10510018 | 10510039 | phastConsElements100way | lod=79 | score:426, srow:38624 |
| 104 | chr1 | 10510060 | 10510064 | phastConsElements100way | lod=13 | score:247, srow:38625 |
| 104 | chr1 | 10510068 | 10510076 | phastConsElements100way | lod=31 | score:333, srow:38626 |
| 104 | chr1 | 10510085 | 10510092 | phastConsElements100way | lod=41 | score:361, srow:38627 |
- 13 featuress
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 104 | cellmeth_recounts | PrEC: 9, LNCaP: 3, DU145: 1 |  |
| 104 | cellmeth_diffreps_PrECvsLNCaP | A=36, B=0, Length=580, Event=Down, log2FC=-Inf, padj=1.48300123068897e-10 |  |
| 104 | cellmeth_diffreps_PrECvsDU145 | A=37, B=1, Length=600, Event=Down, log2FC=-5.21, padj=4.91555057723503e-09 |  |
| 104 | cellexp | id=ENSG00000175279, name=APITD1, PrEC=350, str=36, LNCaP=702, str=470, DU145=267, str=187 |  |
| 104 | cellexp | id=ENSG00000241563, name=CORT, PrEC=6, str=14, LNCaP=8, str=18, DU145=9, str=2 |  |
| 104 | cellexp | id=ENSG00000251503, name=APITD1-CORT, PrEC=0, str=0, LNCaP=0, str=0, DU145=0, str=0 |  |
| 104 | tcgameth | site:cg04113348, B=0.89, L=0.88, H=0.87 |  |
| 104 | tcgameth | site:cg18233538, B=0.92, L=0.91, H=0.91 |  |
| 104 | tcgameth | site:cg24365837, B=0.9, L=0.9, H=0.88 |  |
| 104 | tcgaexp | gene=CORT, entrez=1325, pos=chr1:10509776-10512060(+), B=17, L=21, M=23, H=16 |  |
| 104 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 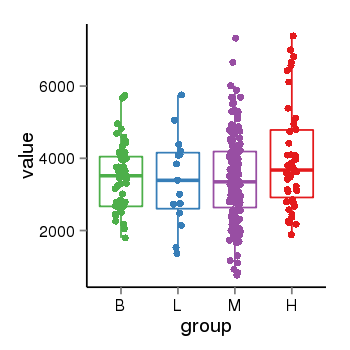 |
| 104 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 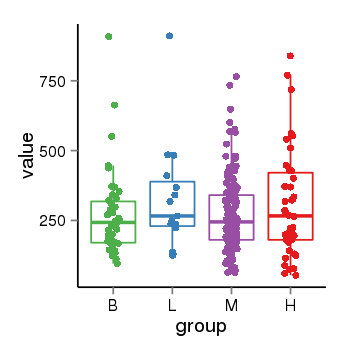 |