DMR 2066 chr4:1211551-1211700 (150bp)
| DMR Stats | |
|---|---|
| Position | chr4:1211551-1211700 (View on UCSC) |
| Width | 150bp |
| Genomic Context | 3' end |
| Nearest Genes | HV535469, CTBP1, HV535487 (0bp away) |
| ANODEV p-value | 0.0186083344466 |
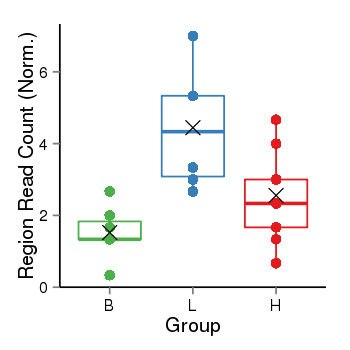
| Frequencies | Benign: 0.0 (0.0%), Low: 4.0 (66.67%), High: 2.0 (22.22 %) |
| Frequent? |
Region Count Boxplot
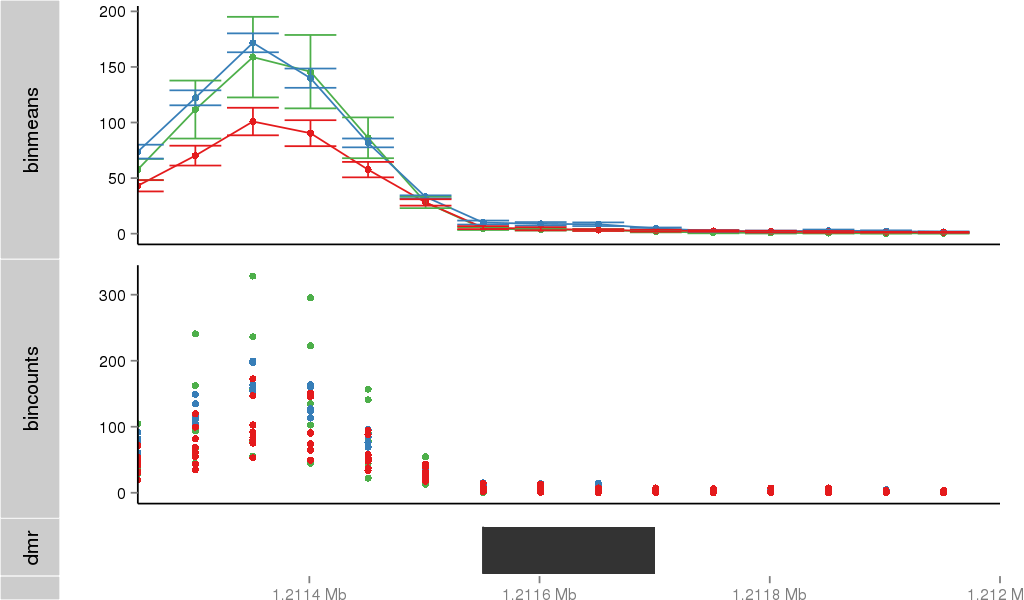
Region Overview
Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2066 | HV535469 | 21280 | uc003gcs.2 | chr4 | 1203908 | 1212379 | + | 150 | 100.0 | 1.24 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (100) | 100.0 |
| 2066 | CTBP1 | 21281 | uc003gcu.1 | chr4 | 1205228 | 1242908 | - | 150 | 100.0 | 0.57 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (100), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0) | 0.0 |
| 2066 | CTBP1 | 21281 | uc003gcv.1 | chr4 | 1205228 | 1242908 | - | 150 | 100.0 | 0.76 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (100), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 2066 | CTBP1 | 21281 | uc003gcw.3 | chr4 | 1205236 | 1242908 | - | 150 | 100.0 | 0.57 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (100), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 2066 | HV535487 | 21280 | uc031sdb.1 | chr4 | 1205822 | 1224816 | + | 150 | 100.0 | 0.76 | 0 | 0.0 | E1 (0), I1 (0), E2 (100) | 0.0 |
| 2066 | CTBP1 | 21281 | uc031sdc.1 | chr4 | 1209779 | 1237041 | - | 150 | 100.0 | 0.57 | 1 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (100), E6 (0) | 0.0 |
- 6 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 2066 | chr4 | 1210194 | 1212193 | cpgShore | — | values(x.gr)[overs$srow, ]:35112 |
- 1 features
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 2066 | cellmeth_recounts | PrEC: 7, LNCaP: 1, DU145: 31 | 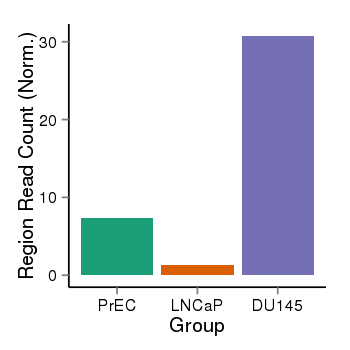 |
| 2066 | cellmeth_diffreps_PrECvsLNCaP | A=456, B=36, Length=1020, Event=Down, log2FC=-3.66, padj=0 | 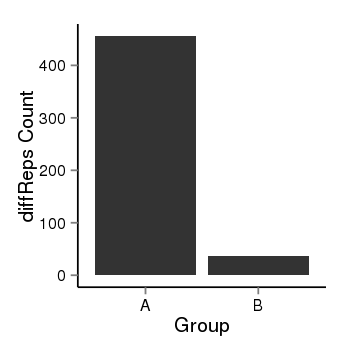 |
| 2066 | cellmeth_diffreps_PrECvsDU145 | A=1219, B=90, Length=2060, Event=Down, log2FC=-3.76, padj=0 | 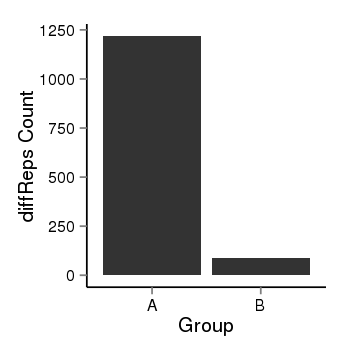 |
| 2066 | cellmeth_diffreps_LNCaPvsDU145 | A=2.14, B=24.32, Length=300, Event=Up, log2FC=3.51, padj=0.000508309456629418 | 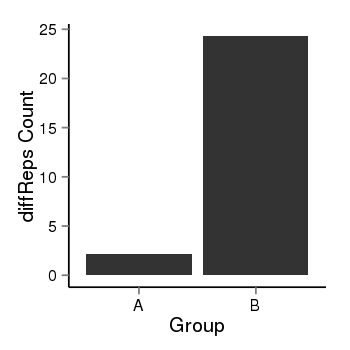 |
| 2066 | cellexp | id=ENSG00000159692, name=CTBP1, PrEC=8103, str=4603, LNCaP=13858, str=8443, DU145=11601, str=9420 | 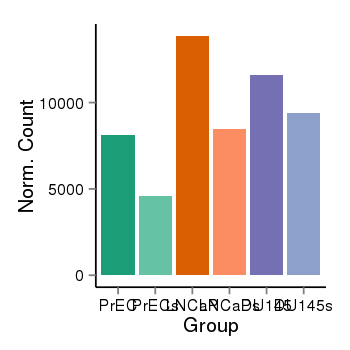 |
| 2066 | tcgaexp | gene=CTBP1, entrez=1487, pos=chr4:1205228-1242908(-), B=7955, L=9441, M=8668, H=8242 | 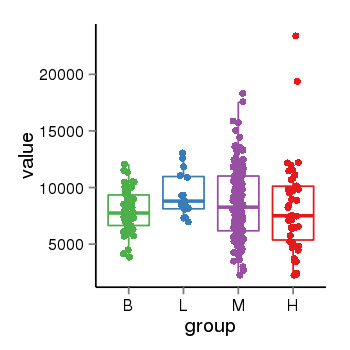 |
| 2066 | tcgaexp | gene=UGT2B10, entrez=7365, pos=chr4:393964-69886113(+), B=12, L=1, M=2, H=8 | 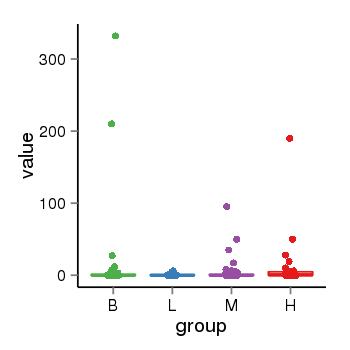 |
| 2066 | tcgaexp | gene=UGT2A3, entrez=79799, pos=chr4:506428-69817509(-), B=1, L=0, M=1, H=1 | 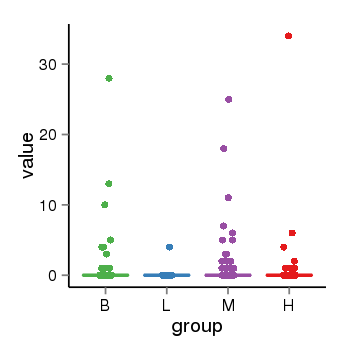 |
| 2066 | tcgaexp | gene=YTHDC1, entrez=91746, pos=chr4:6027-69215824(-), B=4512, L=3976, M=4109, H=4458 | 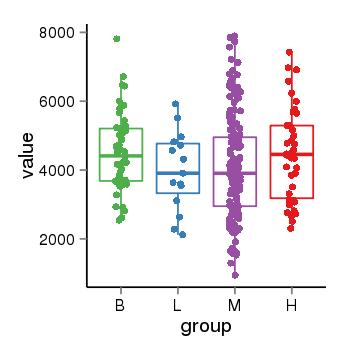 |
| 2066 | tcgaexp | gene=DUX4L5, entrez=653545, pos=chr4:72836-191010142(+), B=0, L=0, M=0, H=0 | 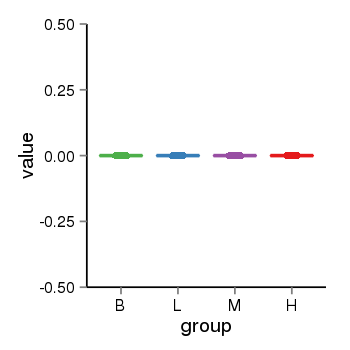 |