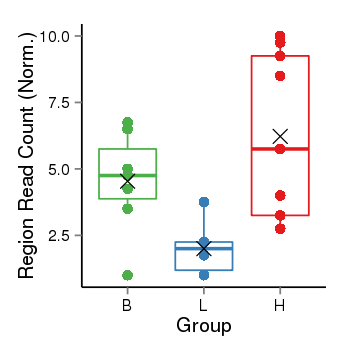
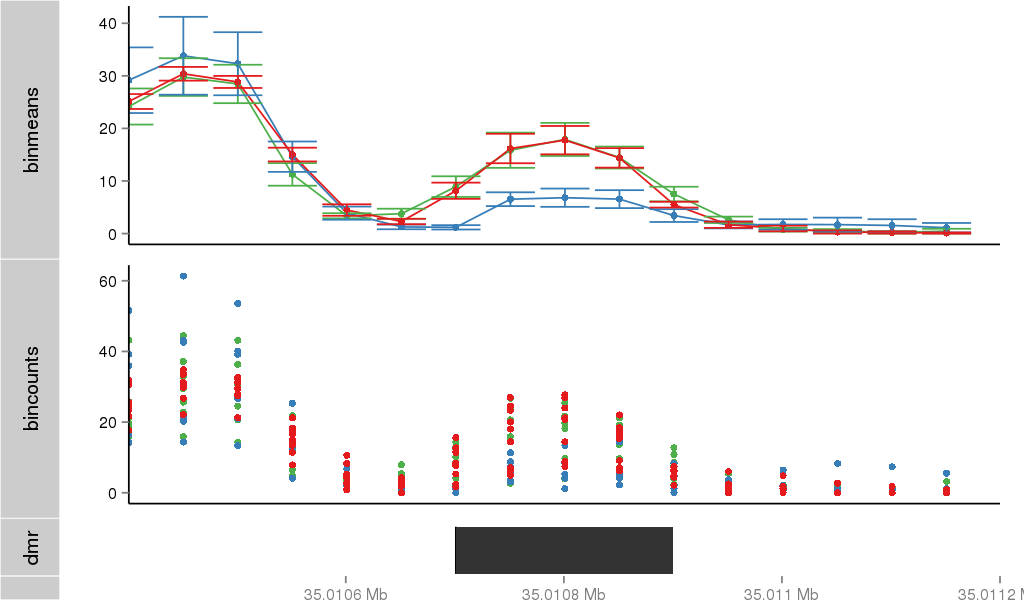
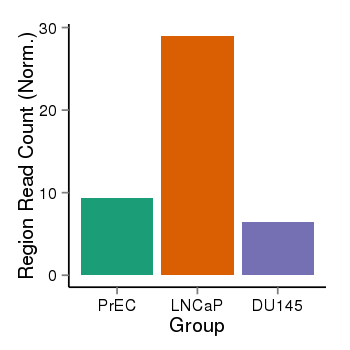| 21800 | cellmeth_recounts | PrEC: 9, LNCaP: 29, DU145: 6 | 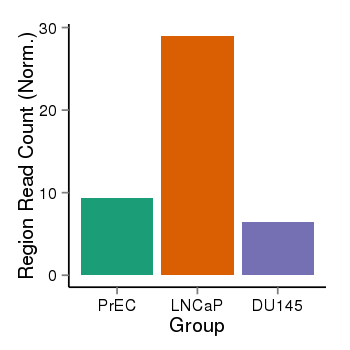 |
| 21800 | cellmeth_diffreps_LNCaPvsDU145 | A=18.17, B=1.87, Length=260, Event=Down, log2FC=-3.28, padj=0.00779054669185256 | 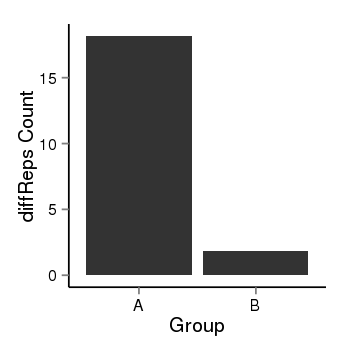 |
| 21800 | tcgaexp | gene=CRHR1, entrez=1394, pos=chr17:954315-43913194(+), B=36, L=21, M=25, H=18 | 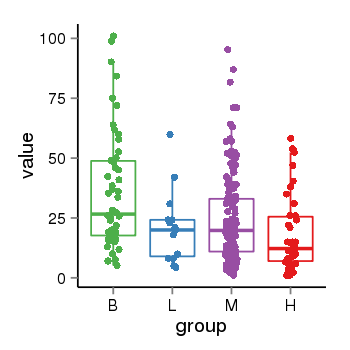 |
| 21800 | tcgaexp | gene=MAPT, entrez=4137, pos=chr17:762281-44105699(+), B=1933, L=2382, M=2487, H=2583 | 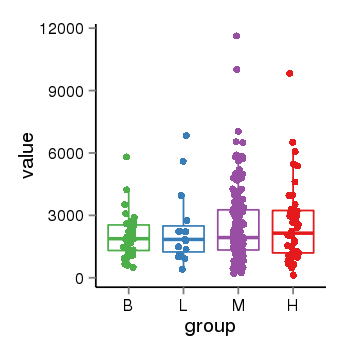 |
| 21800 | tcgaexp | gene=NSF, entrez=4905, pos=chr17:266212-44834828(+), B=3355, L=5678, M=5410, H=5700 | 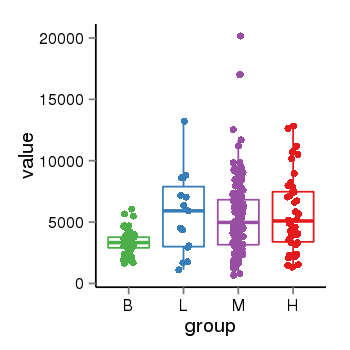 |
| 21800 | tcgaexp | gene=PLEKHM1, entrez=9842, pos=chr17:128328-43568146(-), B=1993, L=1631, M=1589, H=1418 | 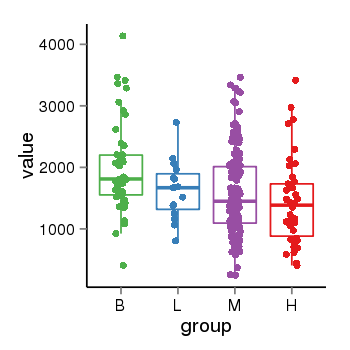 |
| 21800 | tcgaexp | gene=LRRC37A, entrez=9884, pos=chr17:415627-44415160(+), B=924, L=896, M=971, H=940 | 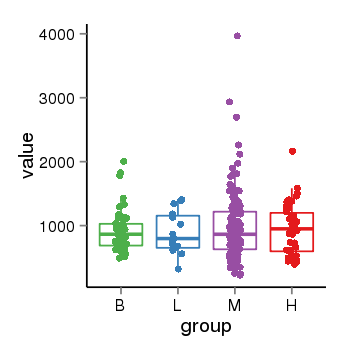 |
| 21800 | tcgaexp | gene=ARL17A, entrez=51326, pos=chr17:195721-44657088(-), B=200, L=195, M=210, H=230 | 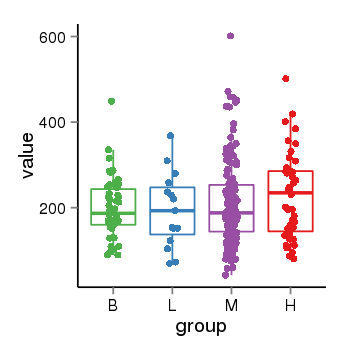 |
| 21800 | tcgaexp | gene=LRRC37A4P, entrez=55073, pos=chr17:197706-43597889(-), B=1015, L=1512, M=1396, H=1450 | 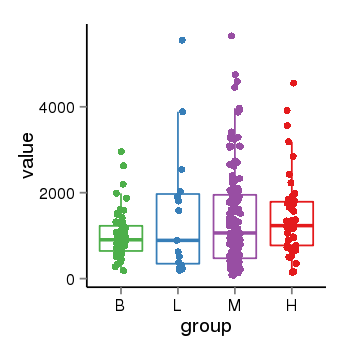 |
| 21800 | tcgaexp | gene=CRHR1-IT1, entrez=147081, pos=chr17:1143726-43723595(+), B=197, L=211, M=215, H=223 | 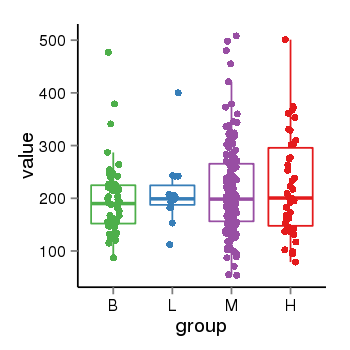 |
| 21800 | tcgaexp | gene=SPPL2C, entrez=162540, pos=chr17:943102-43924438(+), B=0, L=0, M=0, H=0 | 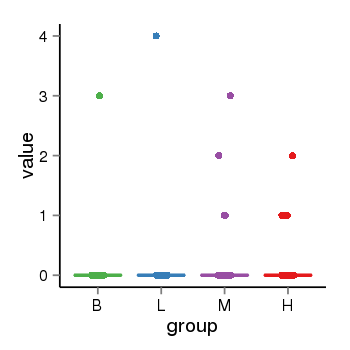 |
| 21800 | tcgaexp | gene=ARHGAP27, entrez=201176, pos=chr17:86404-43511112(-), B=1570, L=1264, M=1273, H=1486 | 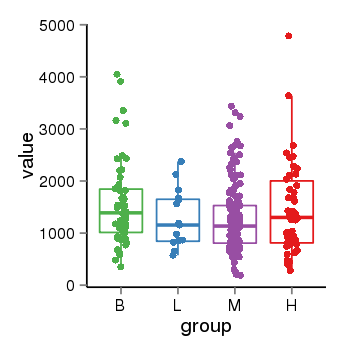 |
| 21800 | tcgaexp | gene=STH, entrez=246744, pos=chr17:790689-44077060(+), B=0, L=0, M=0, H=0 | 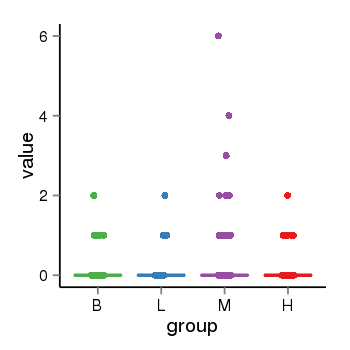 |
| 21800 | tcgaexp | gene=KANSL1, entrez=284058, pos=chr17:563498-44302740(-), B=3141, L=2594, M=2797, H=3017 | 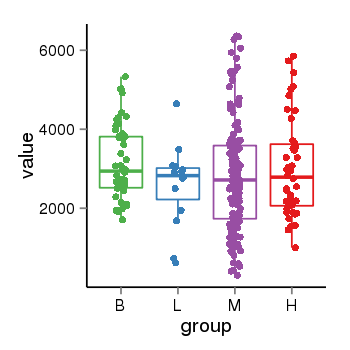 |
| 21800 | tcgaexp | gene=MGC57346, entrez=401884, pos=chr17:1151952-43715329(+), B=601, L=581, M=566, H=598 | 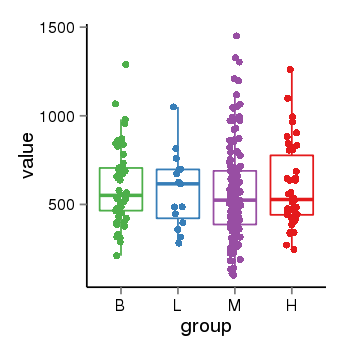 |
| 21800 | tcgaexp | gene=LRRC37A2, entrez=474170, pos=chr17:197706-44633014(+), B=351, L=310, M=361, H=377 | 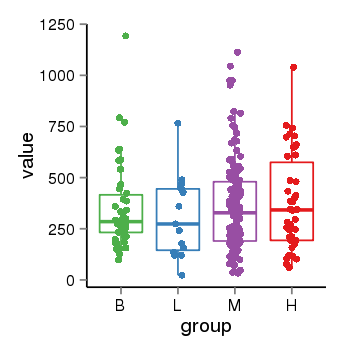 |
| 21800 | tcgaexp | gene=LOC644172, entrez=644172, pos=chr17:1188439-43679748(-), B=84, L=82, M=93, H=90 | 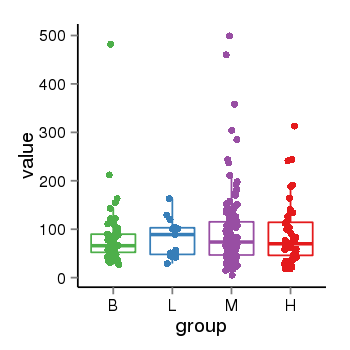 |
| 21800 | tcgaexp | gene=MAPT-AS1, entrez=100128977, pos=chr17:894694-43972879(-), B=1, L=2, M=4, H=2 | 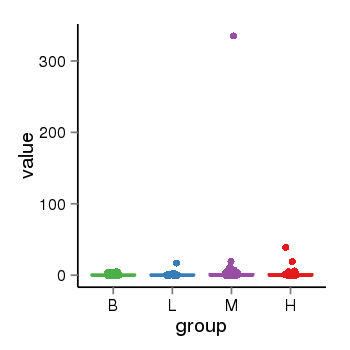 |
| 21800 | tcgaexp | gene=MAPT-IT1, entrez=100130148, pos=chr17:891434-43976164(+), B=5, L=3, M=3, H=4 | 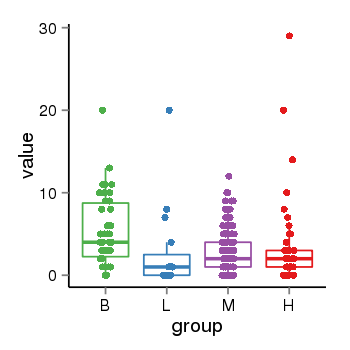 |