DMR 25060 chr1:17201501-17201700 (200bp)
| DMR Stats | |
|---|---|
| Position | chr1:17201501-17201700 (View on UCSC) |
| Width | 200bp |
| Genomic Context | promoter |
| Nearest Genes | CROCC (0bp away) |
| ANODEV p-value | 0.000192092957806 |
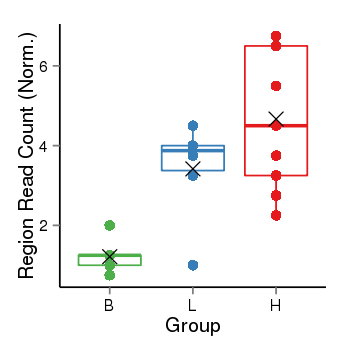
| Frequencies | Benign: 0.0 (0.0%), Low: 5.0 (83.33%), High: 7.0 (77.78 %) |
| Frequent? |
Region Count Boxplot
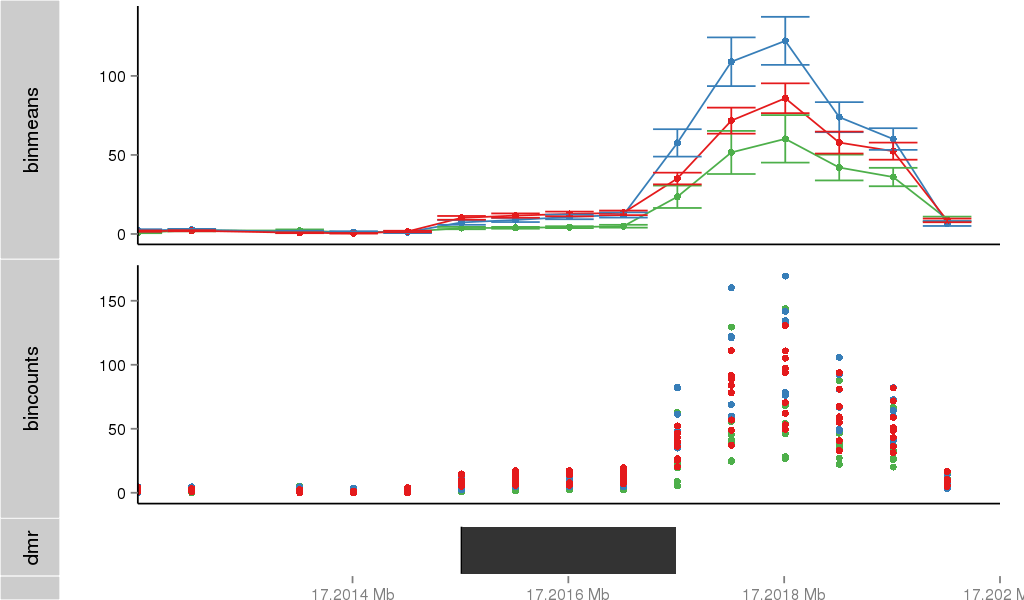
Region Overview
Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 25060 | CROCC | 318 | uc009voy.1 | chr1 | 17066768 | 17267729 | + | 200 | 100.0 | 100.0 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
- 1 genes
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 25060 | chr1 | 17201306 | 17202170 | wgEncodeRegDnaseClusteredV2 | 67 | score:1000, srow:13194 |
| 25060 | chr1 | 17201278 | 17202061 | wgEncodeRegTfbsClusteredV3 | EZH2 | score:251, expCount:1, expNums:36, expScores:251, srow:45233 |
| 25060 | chr1 | 17201328 | 17201997 | wgEncodeRegTfbsClusteredV3 | RBBP5 | score:322, expCount:1, expNums:8, expScores:322, srow:45234 |
| 25060 | chr1 | 17201389 | 17201838 | wgEncodeRegTfbsClusteredV3 | CTCF | score:292, expCount:22, expNums:5,20,37,44,137,595,609,618,636,639,640,641,658,659,661,662,672,673,674,676,677,678, expScores:220,105,136,115,149,184,105,109,123,105,130,90,148,292,237,231,103,225,160,157,127,150, srow:45235 |
| 25060 | chr1 | 17201424 | 17201913 | wgEncodeRegTfbsClusteredV3 | TAF1 | score:162, expCount:1, expNums:157, expScores:162, srow:45236 |
| 25060 | chr1 | 17201494 | 17201733 | wgEncodeRegTfbsClusteredV3 | RAD21 | score:165, expCount:1, expNums:149, expScores:165, srow:45237 |
| 25060 | chr1 | 17200050 | 17202049 | cpgShore | — | values(x.gr)[overs$srow, ]:915 |
- 7 featuress
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 25060 | cellmeth_recounts | PrEC: 5, LNCaP: 3, DU145: 13 | 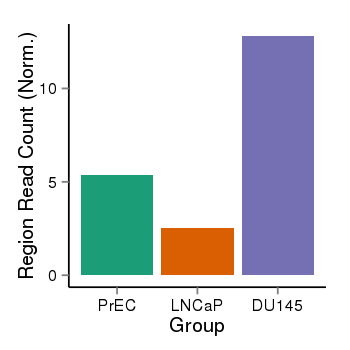 |
| 25060 | cellmeth_diffreps_PrECvsLNCaP | A=104, B=9, Length=460, Event=Down, log2FC=-3.53, padj=0 | 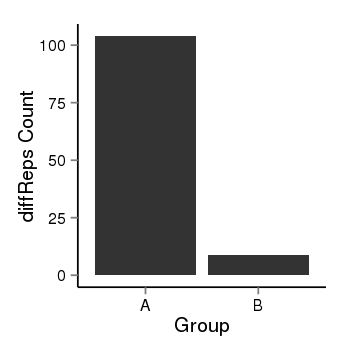 |
| 25060 | cellmeth_diffreps_PrECvsDU145 | A=100, B=19, Length=440, Event=Down, log2FC=-2.4, padj=1.13569037756854e-12 | 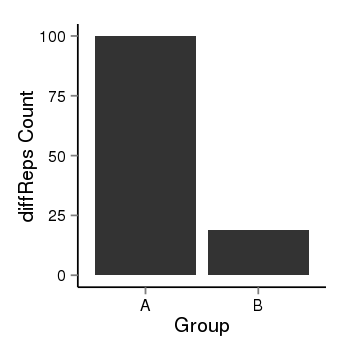 |
| 25060 | cellexp | id=ENSG00000058453, name=CROCC, PrEC=2384, str=844, LNCaP=1171, str=349, DU145=500, str=268 | 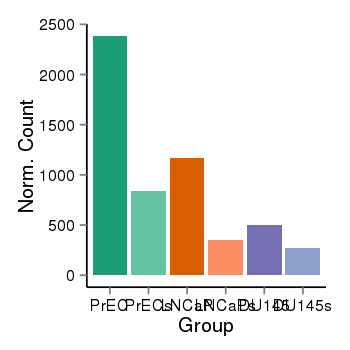 |
| 25060 | tcgaexp | gene=CROCC, entrez=9696, pos=chr1:17066768-17299474(+), B=1096, L=1255, M=1227, H=1094 | 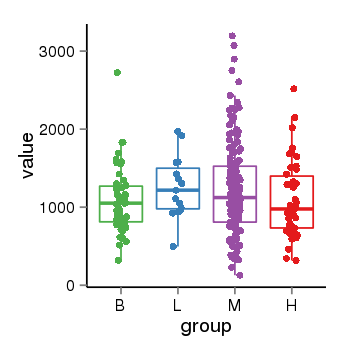 |
| 25060 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 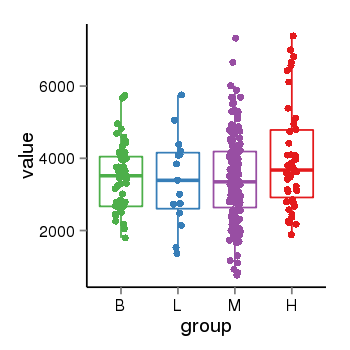 |
| 25060 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 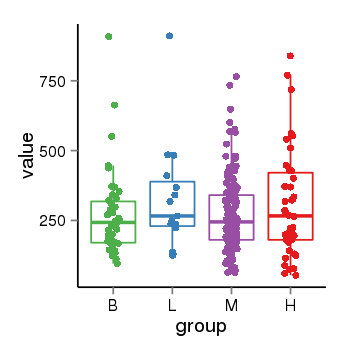 |