DMR 28 chr1:2992151-2992300 (150bp)
| DMR Stats | |
|---|---|
| Position | chr1:2992151-2992300 (View on UCSC) |
| Width | 150bp |
| Genomic Context | intron |
| Nearest Genes | PRDM16 (0bp away) |
| ANODEV p-value | 0.0035646076295 |
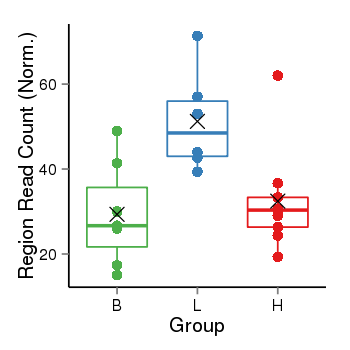
| Frequencies | Benign: 7.0 (100.0%), Low: 6.0 (100.0%), High: 9.0 (100.0 %) |
| Frequent? |
Region Count Boxplot
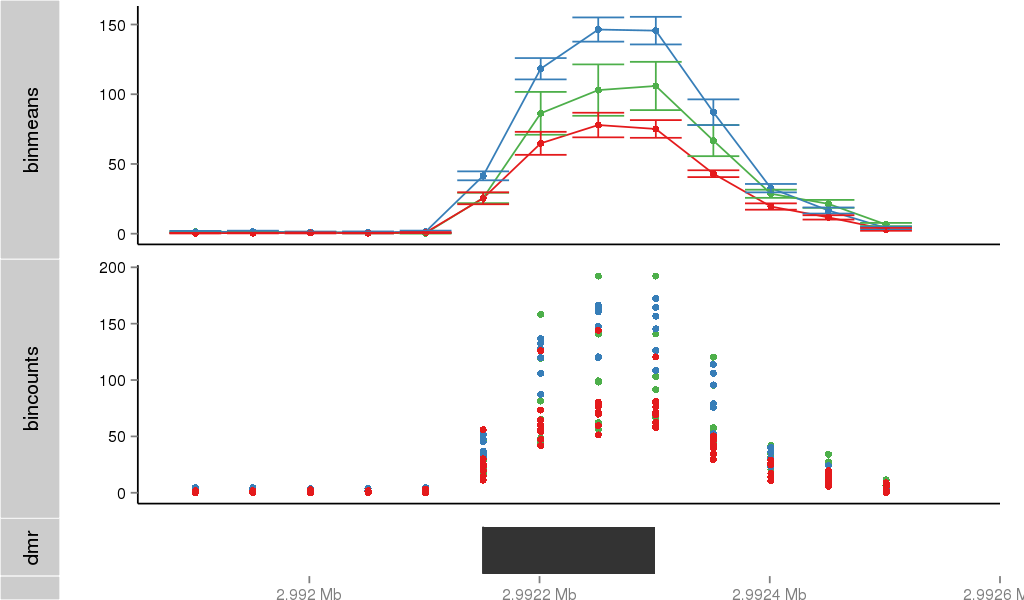
Region Overview
Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 28 | PRDM16 | 117 | uc001akc.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 4.68 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 28 | PRDM16 | 117 | uc001ake.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 0.41 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 28 | PRDM16 | 117 | uc001akf.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 4.68 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 28 | PRDM16 | 117 | uc009vlh.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 0.41 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
- 4 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 28 | chr1 | 2992138 | 2992159 | tfbsConsSites | V$MEF2_03 | score:794, zScore:2.01, srow:5411 |
| 28 | chr1 | 2992140 | 2992154 | tfbsConsSites | V$TATA_01 | score:894, zScore:2.25, srow:5412 |
| 28 | chr1 | 2992155 | 2992170 | tfbsConsSites | V$BRN2_01 | score:837, zScore:2.41, srow:5413 |
| 28 | chr1 | 2992176 | 2992186 | tfbsConsSites | V$POU6F1_01 | score:877, zScore:2.32, srow:5414 |
| 28 | chr1 | 2992180 | 2992187 | tfbsConsSites | V$TBP_01 | score:993, zScore:2.04, srow:5415 |
| 28 | chr1 | 2991794 | 2992205 | phastConsElements100way | lod=4384 | score:824, srow:12767 |
| 28 | chr1 | 2992209 | 2992212 | phastConsElements100way | lod=24 | score:308, srow:12768 |
| 28 | chr1 | 2992215 | 2992232 | phastConsElements100way | lod=80 | score:427, srow:12769 |
| 28 | chr1 | 2992241 | 2992242 | phastConsElements100way | lod=17 | score:274, srow:12770 |
| 28 | chr1 | 2990719 | 2992718 | cpgShore | — | values(x.gr)[overs$srow, ]:383 |
- 10 featuress
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 28 | cellmeth_recounts | PrEC: 146, LNCaP: 77, DU145: 5 | 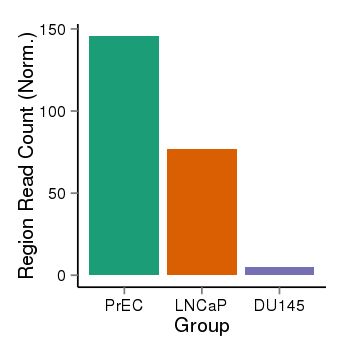 |
| 28 | cellmeth_diffreps_PrECvsLNCaP | A=407, B=95, Length=940, Event=Down, log2FC=-2.1, padj=0 | 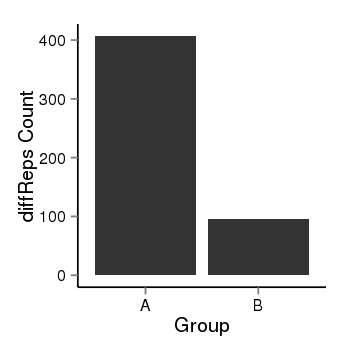 |
| 28 | cellmeth_diffreps_PrECvsDU145 | A=407, B=12, Length=980, Event=Down, log2FC=-5.08, padj=0 | 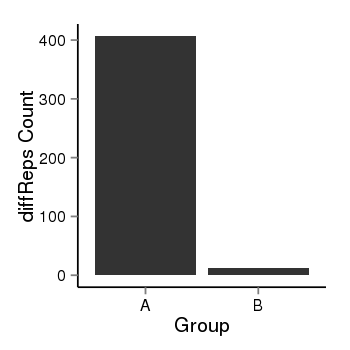 |
| 28 | cellmeth_diffreps_LNCaPvsDU145 | A=73.76, B=5.61, Length=440, Event=Down, log2FC=-3.72, padj=0 | 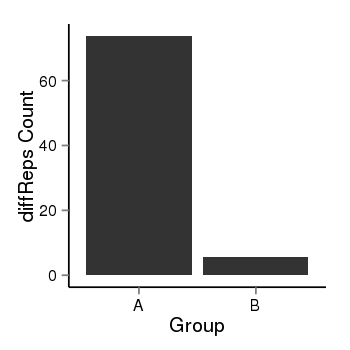 |
| 28 | cellexp | id=ENSG00000142611, name=PRDM16, PrEC=8, str=18, LNCaP=18, str=10, DU145=49, str=42 | 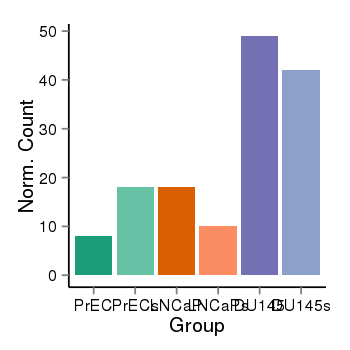 |
| 28 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 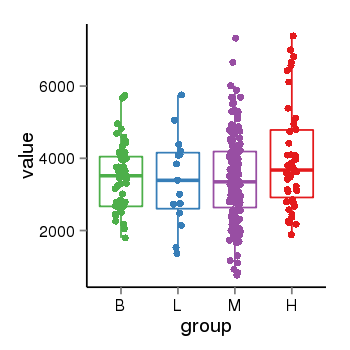 |
| 28 | tcgaexp | gene=PRDM16, entrez=63976, pos=chr1:2985742-3355185(+), B=306, L=40, M=53, H=56 | 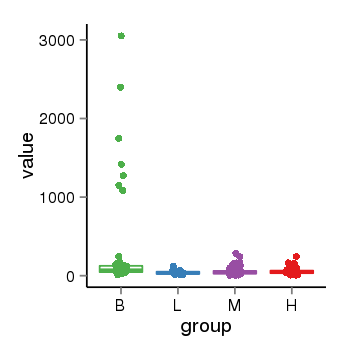 |
| 28 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 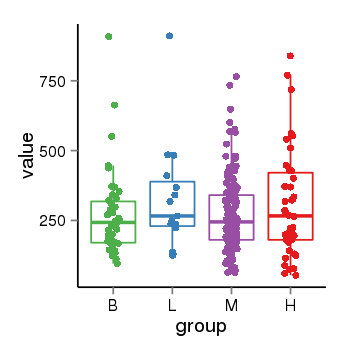 |