DMR 47421 chr1:3072001-3072300 (300bp)
| DMR Stats | |
|---|---|
| Position | chr1:3072001-3072300 (View on UCSC) |
| Width | 300bp |
| Genomic Context | intron |
| Nearest Genes | PRDM16 (0bp away) |
| ANODEV p-value | 0.00421860641917 |
| Frequencies | Benign: 7.0 (100.0%), Low: 6.0 (100.0%), High: 9.0 (100.0 %) |
| Frequent? |
Region Count Boxplot

Region Overview

Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 47421 | PRDM16 | 117 | uc001akc.3 | chr1 | 2985742 | 3355185 | + | 300 | 100.0 | 0.34 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 47421 | PRDM16 | 117 | uc001ake.3 | chr1 | 2985742 | 3355185 | + | 300 | 100.0 | 0.42 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 47421 | PRDM16 | 117 | uc001akf.3 | chr1 | 2985742 | 3355185 | + | 300 | 100.0 | 0.21 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 47421 | PRDM16 | 117 | uc009vlh.3 | chr1 | 2985742 | 3355185 | + | 300 | 100.0 | 0.21 | 0 | 0.0 | E1 (0), I1 (100), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
- 4 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 47421 | chr1 | 3071741 | 3072235 | wgEncodeRegDnaseClusteredV2 | 8 | score:531, srow:2239 |
| 47421 | chr1 | 3072242 | 3072260 | tfbsConsSites | V$OCT1_01 | score:789, zScore:1.78, srow:5766 |
| 47421 | chr1 | 3072246 | 3072259 | tfbsConsSites | V$CEBP_Q2 | score:907, zScore:2.24, srow:5767 |
| 47421 | chr1 | 3072040 | 3072049 | phastConsElements100way | lod=13 | score:247, srow:13166 |
| 47421 | chr1 | 3072052 | 3072063 | phastConsElements100way | lod=14 | score:255, srow:13167 |
| 47421 | chr1 | 3072138 | 3072140 | phastConsElements100way | lod=22 | score:300, srow:13168 |
| 47421 | chr1 | 3072289 | 3072292 | phastConsElements100way | lod=17 | score:274, srow:13169 |
| 47421 | chr1 | 3071900 | 3072239 | cpgIsland | CpG: 27 | length:340, cpgNum:27, gcNum:205, perCpg:15.9, perGc:60.3, obsExp:0.88, srow:243 |
| 47421 | chr1 | 3072240 | 3074239 | cpgShore | — | values(x.gr)[overs$srow, ]:395 |
- 9 featuress
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 47421 | cellmeth_recounts | PrEC: 78, LNCaP: 4, DU145: 13 |  |
| 47421 | cellmeth_diffreps_PrECvsLNCaP | A=171, B=3, Length=820, Event=Down, log2FC=-5.83, padj=0 |  |
| 47421 | cellmeth_diffreps_PrECvsDU145 | A=169, B=14, Length=760, Event=Down, log2FC=-3.59, padj=0 |  |
| 47421 | cellexp | id=ENSG00000142611, name=PRDM16, PrEC=8, str=18, LNCaP=18, str=10, DU145=49, str=42 | 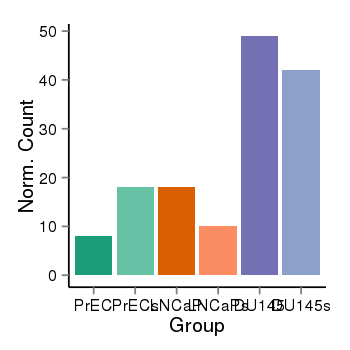 |
| 47421 | tcgameth | site:cg01104489, B=0.74, L=0.63, H=0.62 |  |
| 47421 | tcgameth | site:cg09321238, B=0.75, L=0.69, H=0.65 |  |
| 47421 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 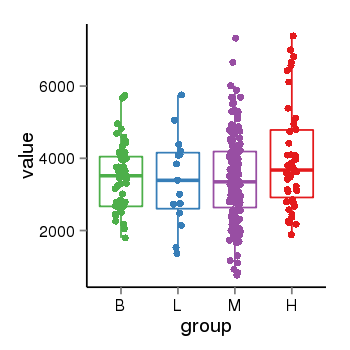 |
| 47421 | tcgaexp | gene=PRDM16, entrez=63976, pos=chr1:2985742-3355185(+), B=306, L=40, M=53, H=56 | 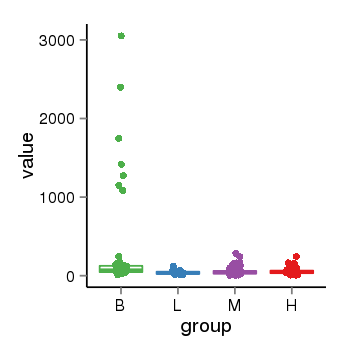 |
| 47421 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 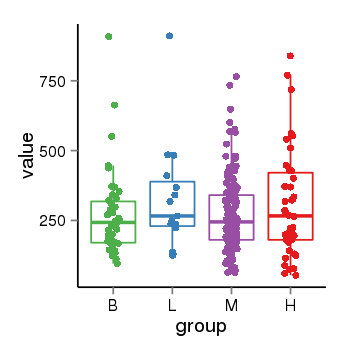 |