DMR 73 chr1:6376051-6376200 (150bp)
| DMR Stats | |
|---|---|
| Position | chr1:6376051-6376200 (View on UCSC) |
| Width | 150bp |
| Genomic Context | intron |
| Nearest Genes | ACOT7 (0bp away) |
| ANODEV p-value | 0.0345740876151 |
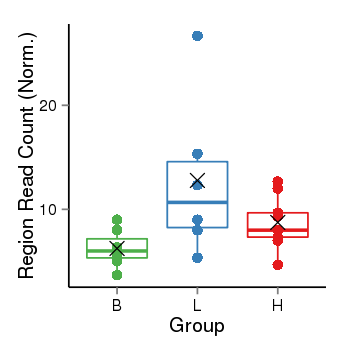
| Frequencies | Benign: 7.0 (100.0%), Low: 6.0 (100.0%), High: 9.0 (100.0 %) |
| Frequent? |
Region Count Boxplot
Region Overview
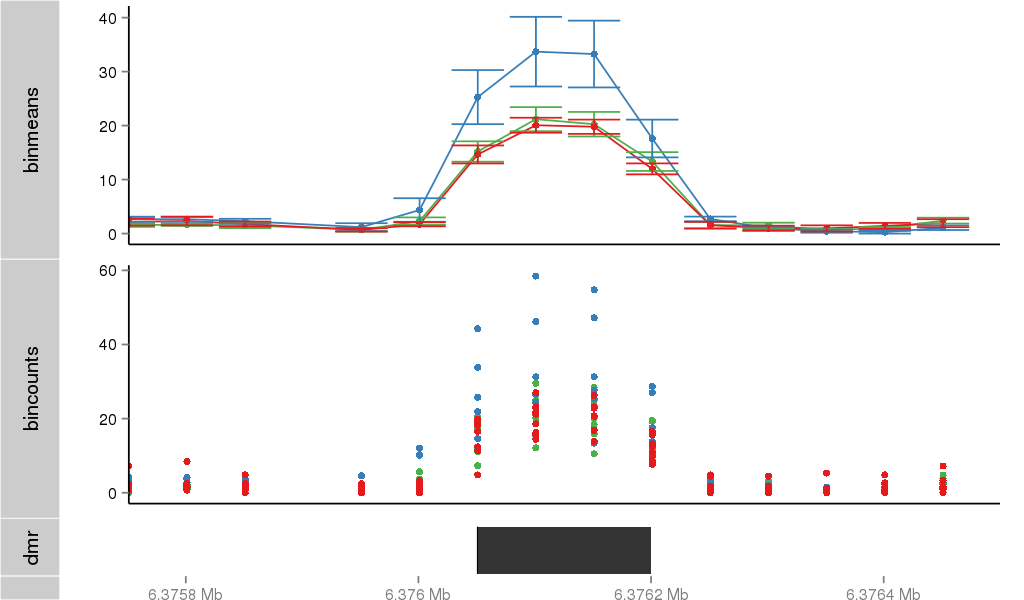
Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 73 | ACOT7 | 149 | uc001amq.3 | chr1 | 6324332 | 6419004 | - | 150 | 100.0 | 0.18 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (100), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 73 | ACOT7 | 149 | uc001amr.3 | chr1 | 6324332 | 6420764 | - | 150 | 100.0 | 0.3 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (100), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 73 | ACOT7 | 149 | uc001ams.3 | chr1 | 6324332 | 6445883 | - | 150 | 100.0 | 0.18 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (100), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 73 | ACOT7 | 149 | uc001amt.3 | chr1 | 6324332 | 6453826 | - | 150 | 100.0 | 0.18 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (100), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0) | 0.0 |
| 73 | ACOT7 | 149 | uc001amu.3 | chr1 | 6324332 | 6453826 | - | 150 | 100.0 | 0.24 | 1 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (0), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (100), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0) | 0.0 |
- 5 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 73 | chr1 | 6376026 | 6376195 | wgEncodeRegDnaseClusteredV2 | 4 | score:223, srow:4922 |
- 1 features
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 73 | cellmeth_recounts | PrEC: 20, LNCaP: 3, DU145: 1 | 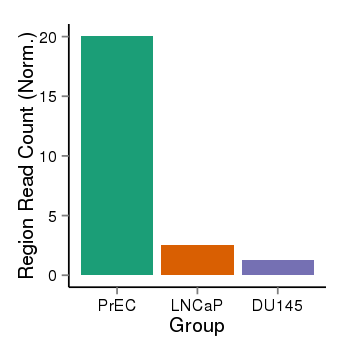 |
| 73 | cellmeth_diffreps_PrECvsLNCaP | A=30, B=2, Length=380, Event=Down, log2FC=-3.91, padj=3.52787813930666e-06 | 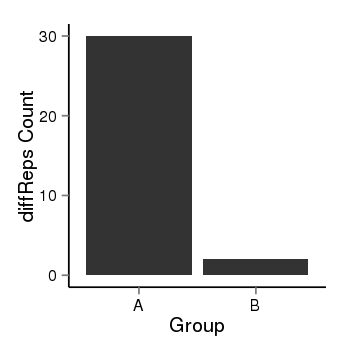 |
| 73 | cellmeth_diffreps_PrECvsDU145 | A=30, B=1, Length=400, Event=Down, log2FC=-4.91, padj=4.59319638008846e-07 | 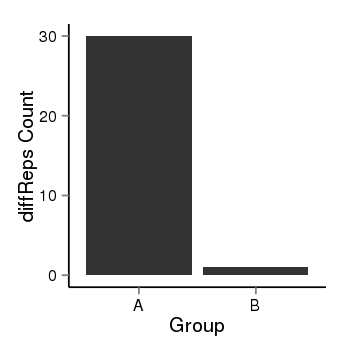 |
| 73 | cellexp | id=ENSG00000097021, name=ACOT7, PrEC=12700, str=8241, LNCaP=2240, str=2875, DU145=2964, str=3293 | 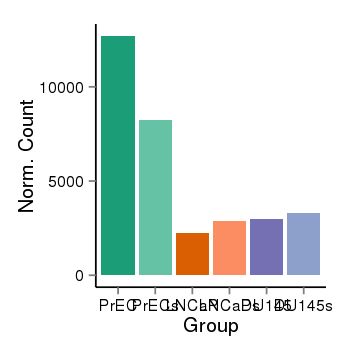 |
| 73 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 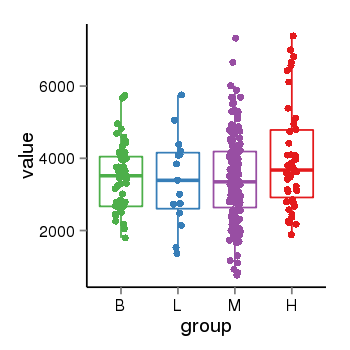 |
| 73 | tcgaexp | gene=ACOT7, entrez=11332, pos=chr1:6324332-6453826(-), B=931, L=1051, M=923, H=1026 | 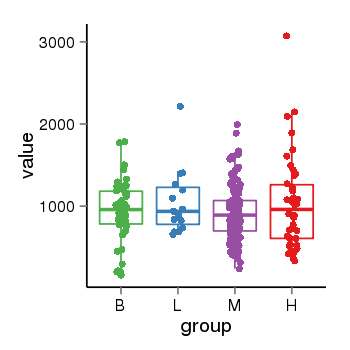 |
| 73 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 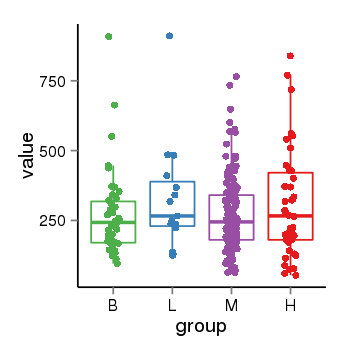 |