DMR 35 chr1:3143551-3143700 (150bp)
| DMR Stats | |
|---|---|
| Position | chr1:3143551-3143700 (View on UCSC) |
| Width | 150bp |
| Genomic Context | intron |
| Nearest Genes | PRDM16 (0bp away) |
| ANODEV p-value | 0.00394834429835 |
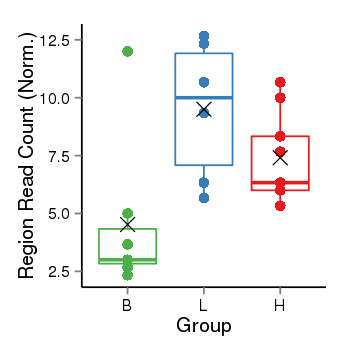
| Frequencies | Benign: 3.0 (42.86%), Low: 6.0 (100.0%), High: 9.0 (100.0 %) |
| Frequent? |
Region Count Boxplot
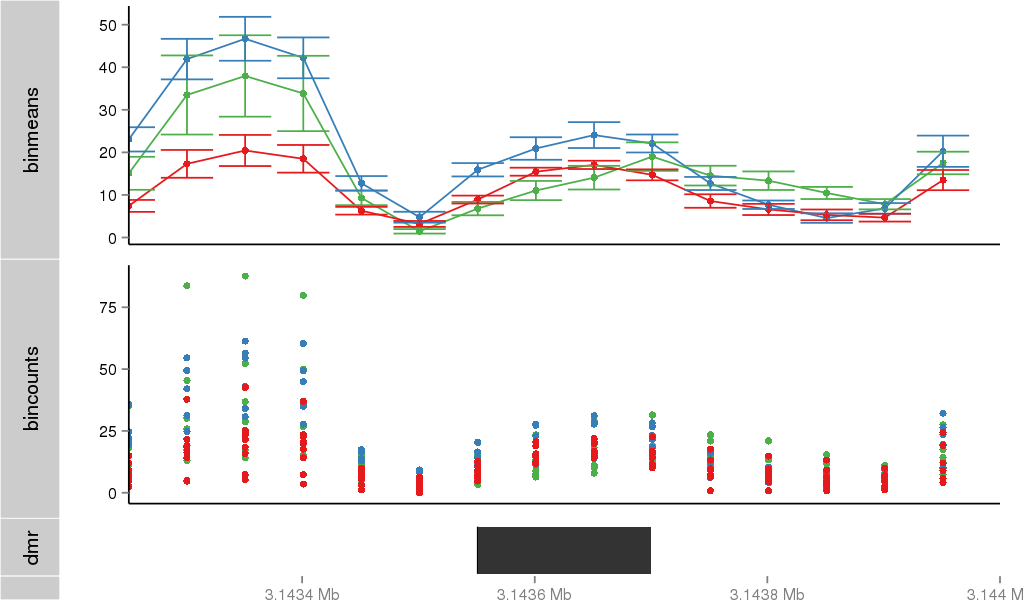
Region Overview
Gene Overlaps
| Dmrid | Gene Symbol | Gene Id | Isoform Id | Isoform Chr | Isoform Start | Isoform End | Isoform Strand | Overlap Bp | Query Overlap Per | Isoform Overlap Per | Noncoding | Promoter Per | Exonintron Per | End Per |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 35 | PRDM16 | 117 | uc001akc.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 1.29 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (100), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 35 | PRDM16 | 117 | uc001ake.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 1.29 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (100), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 35 | PRDM16 | 117 | uc001akf.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 1.29 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (100), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
| 35 | PRDM16 | 117 | uc009vlh.3 | chr1 | 2985742 | 3355185 | + | 150 | 100.0 | 1.51 | 0 | 0.0 | E1 (0), I1 (0), E2 (0), I2 (100), E3 (0), I3 (0), E4 (0), I4 (0), E5 (0), I5 (0), E6 (0), I6 (0), E7 (0), I7 (0), E8 (0), I8 (0), E9 (0), I9 (0), E10 (0), I10 (0), E11 (0), I11 (0), E12 (0), I12 (0), E13 (0), I13 (0), E14 (0), I14 (0), E15 (0), I15 (0), E16 (0), I16 (0), E17 (0) | 0.0 |
- 4 geness
Feature Overlaps
| Dmrid | Feature Chr | Feature Start | Feature End | Set | Name | Fields |
|---|---|---|---|---|---|---|
| 35 | chr1 | 3143568 | 3143573 | tfbsConsSites | V$AML1_01 | score:1000, zScore:1.78, srow:6146 |
| 35 | chr1 | 3143645 | 3143988 | wgEncodeRegTfbsClusteredV3 | GABPA | score:182, expCount:1, expNums:140, expScores:182, srow:8772 |
| 35 | chr1 | 3143668 | 3143907 | wgEncodeRegTfbsClusteredV3 | RAD21 | score:219, expCount:1, expNums:149, expScores:219, srow:8773 |
| 35 | chr1 | 3143625 | 3143634 | phastConsElements100way | lod=14 | score:255, srow:13490 |
| 35 | chr1 | 3143691 | 3143704 | phastConsElements100way | lod=25 | score:312, srow:13491 |
| 35 | chr1 | 3143681 | 3143910 | wgEncodeRegDnaseClusteredV2 | 8 | score:437, srow:2333 |
| 35 | chr1 | 3143575 | 3143620 | phastConsElements100way | lod=123 | score:470, srow:13489 |
| 35 | chr1 | 3143469 | 3143564 | phastConsElements100way | lod=209 | score:522, srow:13488 |
| 35 | chr1 | 3143582 | 3143602 | tfbsConsSites | V$PAX6_01 | score:767, zScore:1.91, srow:6148 |
| 35 | chr1 | 3143635 | 3143654 | tfbsConsSites | V$ARNT_02 | score:770, zScore:1.64, srow:6152 |
| 35 | chr1 | 3143635 | 3143649 | tfbsConsSites | V$E2F_01 | score:777, zScore:2.06, srow:6151 |
| 35 | chr1 | 3143636 | 3143654 | tfbsConsSites | V$PAX2_01 | score:800, zScore:2.35, srow:6153 |
| 35 | chr1 | 3143603 | 3143616 | tfbsConsSites | V$CHX10_01 | score:803, zScore:1.7, srow:6149 |
| 35 | chr1 | 3143575 | 3143592 | tfbsConsSites | V$CEBP_C | score:809, zScore:1.76, srow:6147 |
| 35 | chr1 | 3143690 | 3143703 | tfbsConsSites | V$POU3F2_01 | score:813, zScore:2.09, srow:6154 |
| 35 | chr1 | 3143612 | 3143622 | tfbsConsSites | V$TGIF_01 | score:902, zScore:1.93, srow:6150 |
- 16 featuress
Omics Overlaps
| Dmrid | Omic Set | Data | Plot |
|---|---|---|---|
| 35 | cellmeth_recounts | PrEC: 13, LNCaP: 5, DU145: 8 | 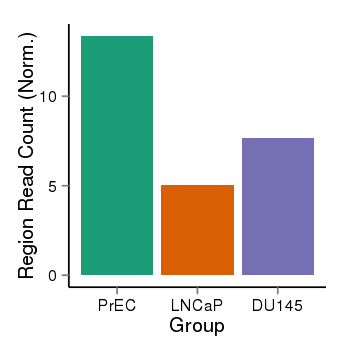 |
| 35 | cellmeth_diffreps_PrECvsLNCaP | A=85, B=4, Length=660, Event=Down, log2FC=-4.41, padj=0 | 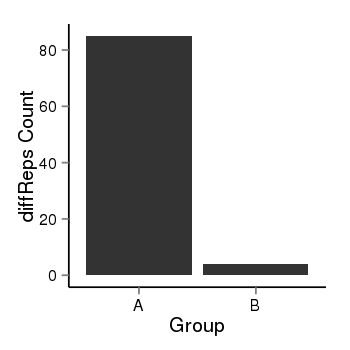 |
| 35 | cellmeth_diffreps_PrECvsDU145 | A=60, B=5, Length=420, Event=Down, log2FC=-3.58, padj=1.78823056245759e-11 | 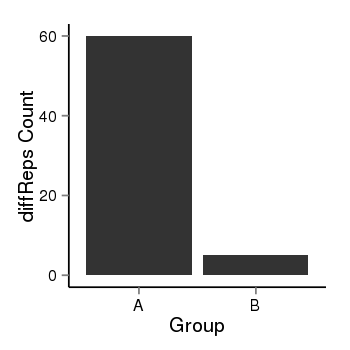 |
| 35 | cellmeth_diffreps_PrECvsDU145 | A=24, B=2, Length=200, Event=Down, log2FC=-3.58, padj=0.000159268381260165 | 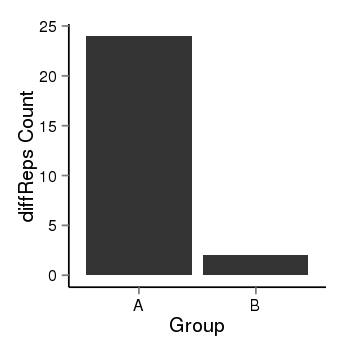 |
| 35 | cellexp | id=ENSG00000142611, name=PRDM16, PrEC=8, str=18, LNCaP=18, str=10, DU145=49, str=42 | 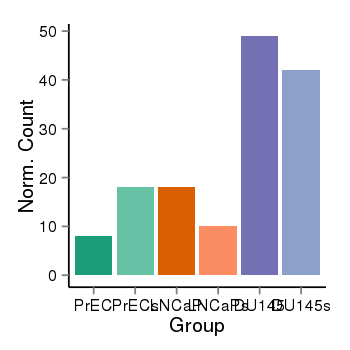 |
| 35 | tcgameth | site:cg23631759, B=0.56, L=0.67, H=0.58 | 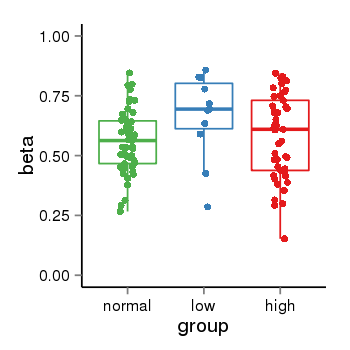 |
| 35 | tcgaexp | gene=SRSF10, entrez=10772, pos=chr1:36273-24306953(-), B=3487, L=3388, M=3432, H=4019 | 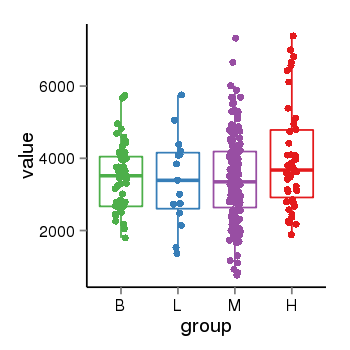 |
| 35 | tcgaexp | gene=PRDM16, entrez=63976, pos=chr1:2985742-3355185(+), B=306, L=40, M=53, H=56 | 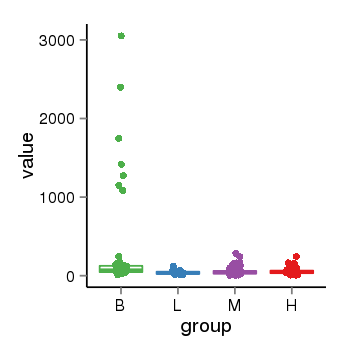 |
| 35 | tcgaexp | gene=LOC100133331, entrez=100133331, pos=chr1:322037-180753553(-), B=268, L=328, M=271, H=305 | 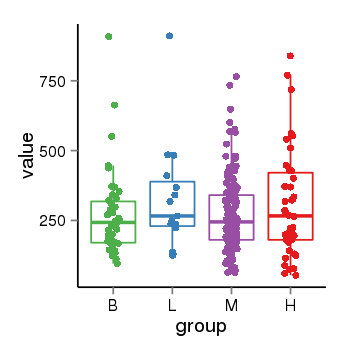 |